Trusted provider of circulating tumor DNA analytics
Blood-based cancer genome profiling
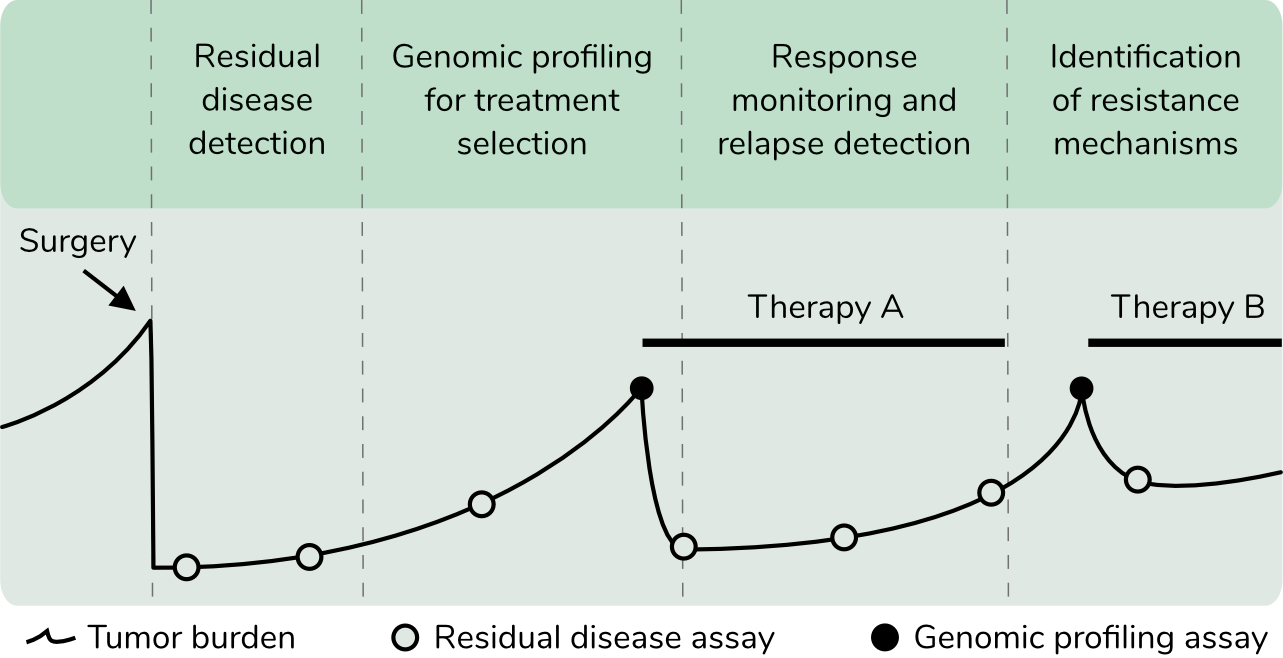
We offer cost-effective and comprehensive solutions for blood-based cancer genome profiling and residual disease detection. Our assays detect somatic mutations, copy number alterations, and chromosomal rearrangements from circulating tumor DNA fragments in blood. The service portfolio also includes customized assays and tumor tissue profiling.
Liquid biopsy analysis
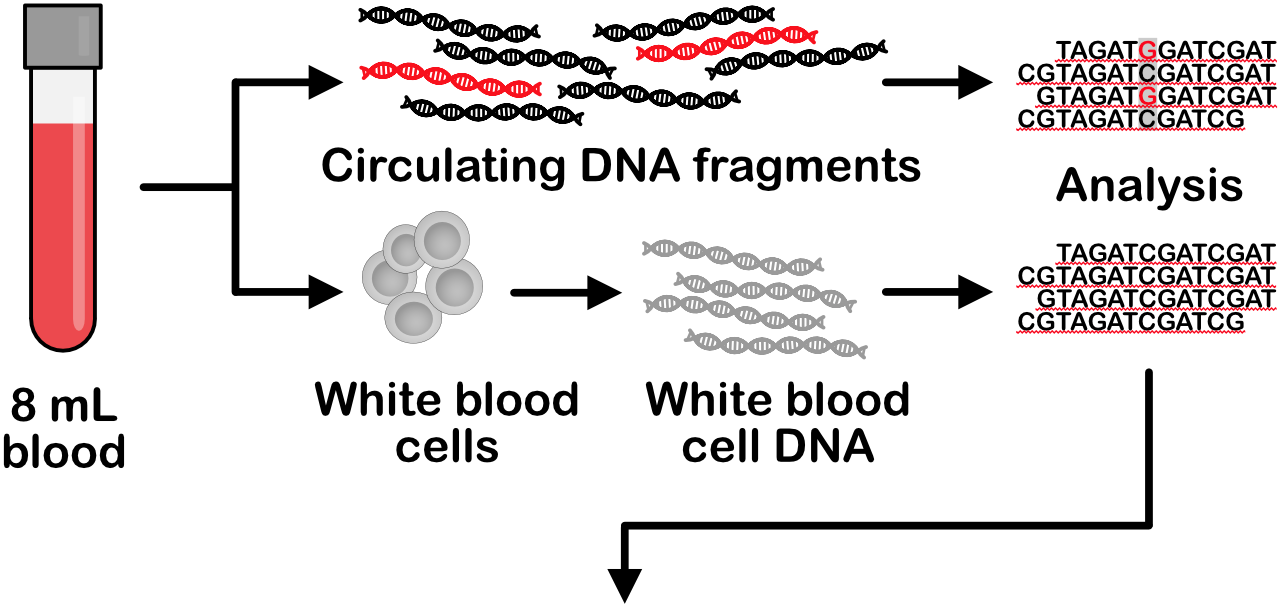
Every person's blood carries billions of DNA fragments released from cells in their body. In a person with cancer, some of these fragments originate from cancer cells. Starting with only 8 mL of patient blood, we extract circulating DNA fragments and analyze them to characterize the cancer cells present in a patient's body. Please see our list of publications for more information.
 View example patient report
View example patient report

